Simple & Fast
Five simple steps with low hands-on time – for results within 24 hours.
Sensitive & Specific
Detect CNVs and selected SNVs.
Clear & High-quality
Clear results and advanced quality checks with Coffalyser.Net.
The power of MLPA in research & diagnostics
SALSA® MLPA® (Multiplex Ligation-dependent Probe Amplification) is the go-to technique for studying gene copy number variations (CNVs) associated with disease. Laboratories worldwide rely on MLPA for the diagnosis and research of genetic disorders and tumours. The power of the technique lies in its versatility: MLPA can be used to detect copy number changes in anything from complete chromosomes to single exons. MLPA is also used to detect DNA methylation changes (MS-MLPA) and is sensitive enough to discriminate aberrations in disease-causing genes from highly similar pseudogenes.
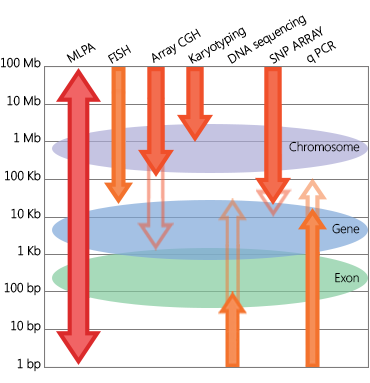
Why MLPA?
CNVs play a role in hundreds of genetic disorders, and MLPA assays exist for even the rarest of them. Laboratories around the world use MLPA to ensure that deletions and duplications are adequately recognised. For disorders where CNVs make up the majority of mutations, MLPA is used as a first-line test. The same applies to disorders in which genetic analysis is complicated by pseudogenes: here, MLPA is used for its exceptional sensitivity to discriminate closely related sequences. In oncogenomics, MLPA is used to study somatic CNVs to optimise tumour profiling.

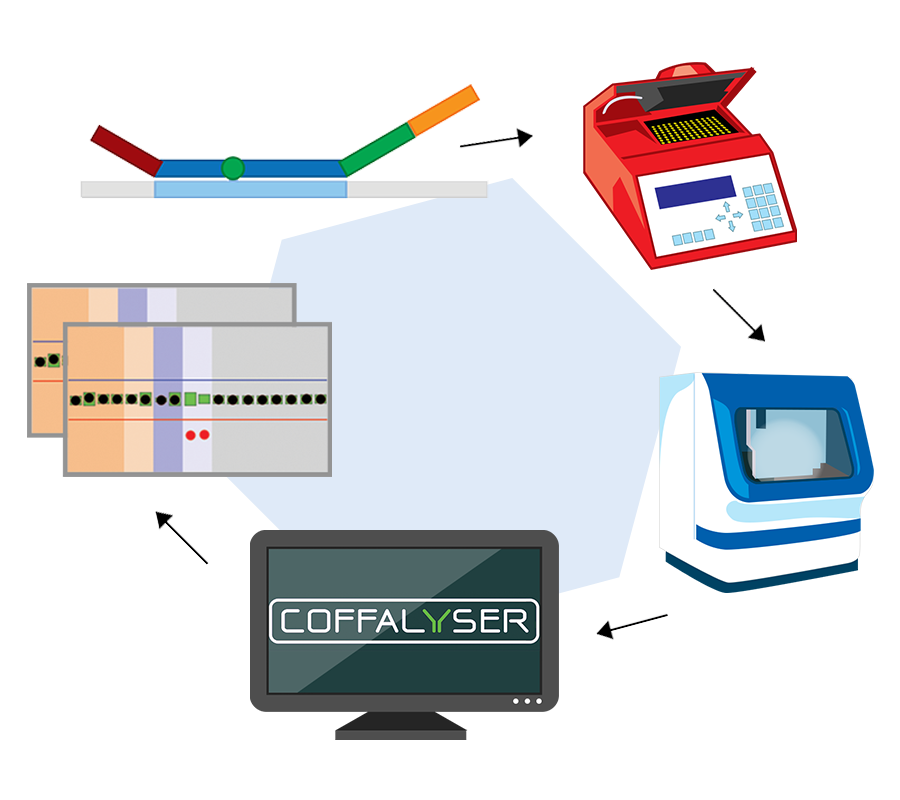
How does MLPA work?
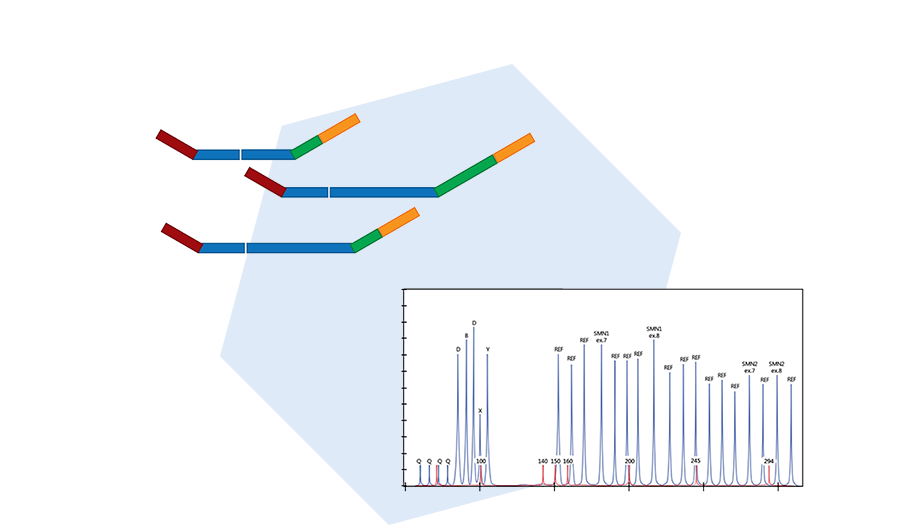
MLPA is a simple, multiplex PCR technique that uses a single primer pair to amplify up to 60 probes, each with a unique genomic target and length. PCR amplicons are fluorescently labelled and separated and quantified by capillary electrophoresis. By comparing the resulting peak pattern of a sample to those of a set of reference samples, the number of genomic targets present in the sample of interest can be determined.
New to MLPA?
If you are new to MLPA, you may want to have a look at our guide to getting started. In it, we have listed all the best resources for beginners in a stepwise fashion. Via videos, e-learning modules and background explanations you will learn more about the technique, the practical execution of the protocol, how to perform data analysis and quality control, and where to find more detailed information.
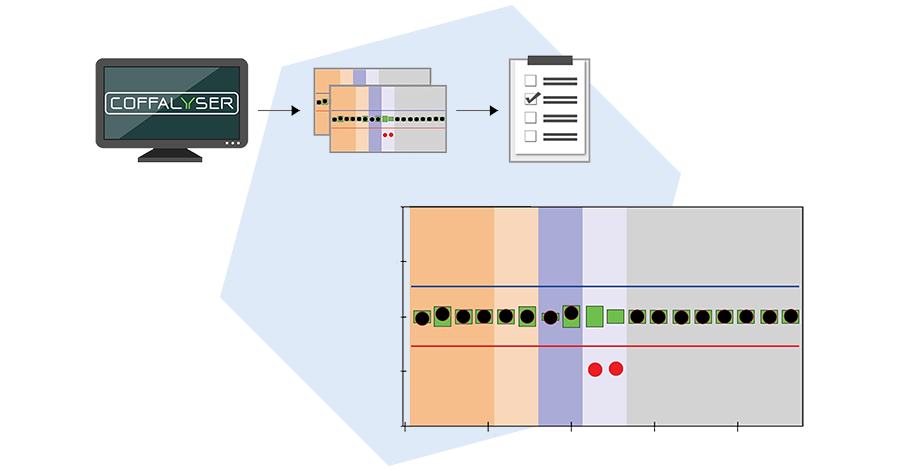
Coffalyser.Net: MLPA analysis software
Coffalyser.Net™ is free MLPA analysis software made and supported by MRC Holland. Coffalyser.Net can directly import raw data files, and performs advanced quality checks to increase the robustness of your results. Analysis sheets for each product can be retrieved directly from our servers, ensuring you always have access to the latest versions.
Can We Help You?
Contact our support team.



